Main Page
From Bioinformatics Software
Jump to navigationJump to searchWelcome to the Bioinformatics Software and Resources page.
Software
Key: P = phylogenetics, S = statistics, B = biogeography, V = visualization, G = genomics, M = metagenomics, L = lateral genetic transfer, A = sequence alignment
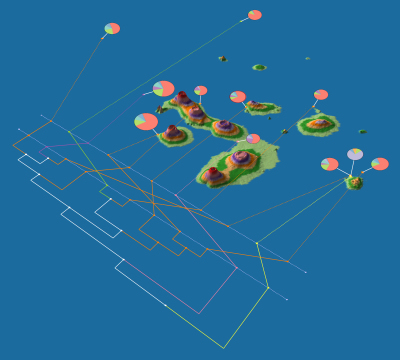
- GenGIS: an application that allows users to combine digital map data with information about biological sequences collected from the environment. GenGIS provides a 3D graphical interface in which the user can navigate and explore the data, as well as a Python interface that allows easy scripting of statistical analyses using the Rpy libraries. (PSBVM)
- Radié: a tool that allows characters to be visualized against the background of a phylogenetic tree. The software includes several different visual and numeric representations of the ‘convexity’ of a given character, in other words the extent to which different character traits form distinct groups within the tree. (PV)
- STAMP: a software package for analyzing metagenomic profiles that promotes ‘best practices’ in choosing appropriate statistical techniques and reporting results. (SMV)
- EvolSimulator: a simulation test bed for hypotheses of genome evolution. (PL)
- EEEP (Efficient Evaluation of Edit Paths): software to infer putative pathways of lateral genetic transfer by comparing gene trees against a rooted reference tree (PL)
- WOOF: a tool designed to rigourously apply the principle of visual alignment validation. (SA)
- GANN: a machine learning method designed with the complexities of transcriptional regulation in mind.
- VAREB?
Web Services
- SeqMonitor
- MOA (to be added)
- Visual MOA (to be added)
- MANUEL
Datasets
- Datasets used in GenGIS (Parks et al., Genome Research 2009)
- STAMP datasets
- Simulated data for 'The impact of reticulate evolution on genome phylogeny' (Beiko et al., Systematic Biology 2008)
- LGT in 144 genomes