STAMP
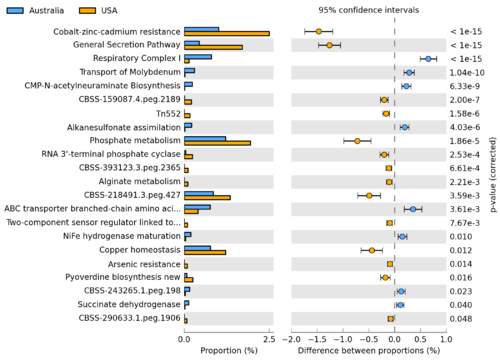
STAMP is a software package for analyzing taxonomic or metabolic profiles that promotes ‘best practices’ in choosing appropriate statistical techniques and reporting results. Statistical hypothesis tests for pairs of samples or groups of samples is support along with a wide range of exploratory plots. STAMP encourages the use of effect sizes and confidence intervals in assessing biological importance. A user friendly, graphical interface permits easy exploration of statistical results and generation of publication quality plots for inferring the biological relevance of features in a metagenomic profile. STAMP is open source, extensible via a plugin framework, and available for all major platforms.
Announcements
- December 5, 2014: STAMP v2.0.9 released. Improved handling of missing sample metadata. Rows and columns in heatmaps can now be sorted alphabetically.
- August 5, 2014: STAMP v2.0.8 released. Explicitly checks that STAMP profiles form a strict hierarchy. Permits generation of larger heatmaps.
- July 29, 2014: STAMP v2.0.7 released. Enhanced heat map plots. Improved support for numpy v1.8.1+.
- July 7, 2014: STAMP v2.0.6 released. STAMP v2.0.5 erroneously contained the BIOM v1 library instead of BIOM v2.
- June 29, 2014: STAMP v2.0.5 released. Improved support for BIOM v2, updated User's Guide, improved handling of large datasets with heat map plots.
- June 18, 2014: STAMP v2.0.4 released. Migrated to BIOM v2 format.
- Previous announcements
Documentation
- Quick installation instructions (Microsoft Windows, Linux, Apple's Mac OS X)
- User's Guide
- Google Group
- FAQs
- Version history
Downloads
Please uninstall previous versions of STAMP before installing a new release.
- STAMP v2.0.9 setup package for Microsoft Windows (~42MB)
- Linux and OS X users can follow the instructions to install from source
- STAMP GitHub Repository
- Previous versions
Additional Scripts
- checkHierarchy.py: identify entries within a STAMP profile that are not strictly hierarchical
Examples
Citing STAMP
If you use STAMP v2+ in your research, we appreciate you citing:
- Parks DH, Tyson GW, Hugenholtz P, Beiko RG. (2014). STAMP: Statistical analysis of taxonomic and functional profiles. Bioinformatics, 30, 3123-3124.
STAMP v1 manuscript:
- Parks DH and Beiko RG. (2010). Identifying biologically relevant differences between metagenomic communities. Bioinformatics, 26, 715-721.
Contact Information
STAMP is in active development and we are interested in discussing all potential applications of this software. We encourage you to send us suggestions for new features. Suggestions, comments, and bug reports can be sent to Rob Beiko (beiko [at] cs.dal.ca). If reporting a bug, please provide as much information as possible and a simplified version of the data set which causes the bug. This will allow us to quickly resolve the issue.
Funding
The development and deployment of STAMP has been supported by several organizations:
- Genome Atlantic
- The Dalhousie Centre for Comparative Genomics and Evolutionary Bioinformatics, and the Tula Foundation
- Killam Trust
- The Natural Sciences and Engineering Research Council of Canada
- The Dalhousie Faculty of Computer Science